
Towards models of infection transmission dynamics
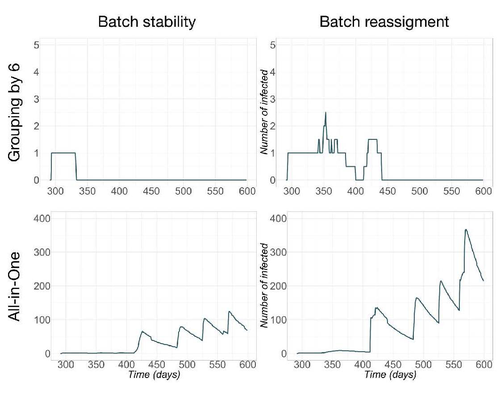
Pig herd management and infection transmission dynamics: a challenge for modellers.
Abstract
Recommendation: posted 09 February 2024, validated 09 February 2024
Letourneau Montminy, M.-P. (2024) Towards models of infection transmission dynamics. Peer Community in Animal Science, 100198. https://doi.org/10.24072/pci.animsci.100198
Recommendation
Epidemics such as PRRSv-like virus in pig farms has tremendous impact on the competitiveness of swine production. However, its control requires an understanding of the complex interaction between pathogen transmission, disease impact, population dynamics and management. By using mechanistic epidemiological modelling, Sicard et al. (2023) open up a very interesting field of possibilities. This article describes work aimed at assessing the consequences of infections, taking into account the interaction between clinical outcomes and population dynamics. This study shows how this interaction can influence transmission dynamics at the herd level. It highlights the need to further explore this direction, integrating both disease impacts in breeding practices and structural changes in population dynamics, such as pig crossbreeding and grouping methodologies.
The provision of a new tool making it possible to model herd management practices and the transmission of a virus, such as PRRS, in time and space is a major contribution to understanding the dynamics of this category of diseases. It opens up the possibility of being able to represent specific herd structures and evaluate transmission dynamics using real data. This work improves our understanding of disease spread across herds, taking into account herd management.
Reference
Sicard V, Picault S, Andraud M (2023) Pig herd management and infection transmission dynamics: a challenge for modellers. bioRxiv, 2023.05.17.541128. ver. 2 peer-reviewed and recommended by Peer Community in Animal Science. https://doi.org/10.1101/2023.05.17.541128
The recommender in charge of the evaluation of the article and the reviewers declared that they have no conflict of interest (as defined in the code of conduct of PCI) with the authors or with the content of the article. The authors declared that they comply with the PCI rule of having no financial conflicts of interest in relation to the content of the article.
INRAE (Animal Health Division), French Region Pays de la Loire
Evaluation round #1
DOI or URL of the preprint: https://doi.org/10.1101/2023.05.17.541128
Version of the preprint: 1
Author's Reply, 18 Dec 2023
Decision by Marie-Pierre Letourneau Montminy, posted 20 Sep 2023, validated 21 Sep 2023
Dear Vianney Sicard,
Thank you for submitting your manuscript to PCI Animal Science.
Your submission has been reviewed by two independent reviewers who found merit in your work. They provided feedback to improve the manuscript.
I invite you to submit a revised version of your manuscript addressing the comments of the reviewers.
Best regards
Marie-Pierre Letourneau Montminy
Reviewed by Gustavo Machado, 30 Jul 2023
This manuscript describes a novel model simulation tool that can capture multiple disease dynamics levels. This work enhances our understanding of whole herd disease spread, taking into consideration herd management.
I have noticed some spacing and a lack of capital letters in the abstract; please reread it you will find it
Line 31: Please add a citation to the end of the sentence
Line 36: Also, a citation after the mention of AI would be helpful
I would also suggest you remove the many ..etc. from your examples it is not necessary, you could use, e.g., to represent that those are examples but not limited to that list
Line 80: When you describe the levels in which you say fine-grained level, it would be useful if you could add a figure here to represent all levels you have included in your model
Line 105: For the physiological states you have fixed it, is it possible to comment on the possibility of varying it around a mean value, given that production is never perfectly dated?
Line 113: It would help to have a diagram showing how rooms were divided to see where the direct and indirect transmission forces act.
While Figures 4 and 5 are useful for EMULATION user, adding a diagram with it would make this more friendly for no-EMULATION users.
In your pen-level network, nodes were metapopulation or agents?
For all scenarios, the introduction of the infected gilt was at the same pen? If not, could up please describe, if no, where was this pen located in the barn?
Line 238: You say several studies and cite only one; please cite more
Line 240: Also missing appropriate citations
Line 264: Also missing citation
https://doi.org/10.24072/pci.animsci.100198.rev11
Reviewed by anonymous reviewer 1, 06 Aug 2023
This is a very interesting and well written article. The construction of a modeling framework making it possible to model herd management practices and the transmission of a virus, such as PRRS, in time and space is a major contribution to understanding the dynamics of this category of disease.
I think that the choices of parameters made in M&M must be better explained and discussed in the Discussion section. Overall, the M&M and discussion parts need to be improved.
Make better use of the Figures would be profitable, in particular the diagrams by adding the functions and the steps to which they are applied (culling state machine, inseminationStatus…) From my point of view, Figures including coding do not necessarily provide information for understanding (or they are not sufficiently exploited). I think they could be added as additional material.
Some of my remarks may be related to misunderstandings on my part., so feel free to disagree.
Introduction
14- The
19 – Avoid acronyms in the abstract or define SEIR.
30 - Develop shortly the concept of multi-level agent-based modelling or add a reference
34-37 – This sentence is not very understandable for me, especially the contributions of “Artificial Intelligence methods”. It might be better to rephrase.
38-40 – Based on EMULSION software ?
43-46 – Is this the purpose of this study or is this a previous project? If it is the purpose of this study, it would be better to add this sentence to the end of the section.
52-55 – MLABS are an evolution of ABS models ? These models are included in EMULSION framework? The relationship between these approaches is not clear for me.
67 – Specify that in this article the scenarios tested are on the management policy of breeding sector (reassignment / sow grouping).
Materials and Methods
70-73 – Perhaps these 2 sentences are more suitable as an introduction for the presentation of EMULSION?
74 – Population dynamics section ?
75 – It would be better to rephrase, by avoiding use of ().
101 – The information is already written line 97.
105 – insemination (24 days) à 34 days (Figure 2).
Figure 2 it might be useful to represent the culling stage or the impact on the cycle of insemination failure. Figures 2 and 3 could be combined into one.
Figure 4 is not really exploited in the text. Is it relevant? Or perhaps add as supplementary material.
121 - The tested approach does not consider the possibility of performing adoptions between litters in the same “batch” (or between batches ?)
122 – 18 pens per batch and per organizational level ? (insemination / gestating / farrowing ?). The relationship between 29 sows at initialization and the organizational level of 18 pens is not clear for me. It could be interesting to add this information on Figure 2.
134 – Remove “.”
140 - How does the model arbitrate between the replacement of a culled sow and a dead sow? Are gilts used primarily to replace dead sows and then culled sows? More generally, how is the sow mortality rate simulated/managed?
Figure 5 is not cited in the text.
149 – “physiologicalStep” in italic ?
150 – inseminate status is different than inseminationStatus ? and X days ? gestation control is not performed at fixed duration after insemination ?
151 – How “proba_failure_ins” is determined ?
159 – This information is not represented in Figure 3.
157 – 160 It would be necessary to further explain the 2 machine states : health_state and maternal_immunity or to refer to a previous article.
Table 1 – “Duration in M” is related to duration of state M ?
163 – Add a sentence to explain the choice of the 3 other parameters.
166-167 – A Figure to illustrate could be useful.
168 – Is it related to the “nodes” cited in legend of Figure 5 ?
186 – And abortion ?
187 – table 2.4 ? – the rate occasional contact is related to batch reassignment policy ?
Table 2 – In results section, you are not using the scenario numbers but rather the Bind parameter. Add for each scenario tested, the corresponding parameter.
Results
Results Section – Named the corresponding scenarios
195 – One room per batch is related to “All-in-One grouped sows” ? it is not ldescribed in the 4 scenarios.
Figure 6 – Named the corresponding scenario number – For the time scale, explain that it is the number of days since the beginning of the simulation (or start at 0 = day of infection to be consistent with the text).
209 – 42 days is X on line 150 ?
Discussion
I think the Discussion part needs to be improved. In my view, the choices of model parameters are not discussed clearly enough.
It might be interesting to discuss the robustness of the modeling approach (related to RFF), and in particular its sensitivity to the various parameters tested.
I think it would be interesting to discuss more about the contributions of this model compared to the one tested on Influenza A.
230 – Maybe, discuss more the contributions of AI based approach to model interplay between herd organization and disease dynamics (or refer to previous article).
261 - I am not an epidemiologist and therefore it may be linked to my lack of understanding, but from my point of view the "indirect contact rate" parameter is not sufficiently explained in the scenarios.
262 - Are the number of infected animals and the transmission dynamics presented representative of the information available in the state of the art?
https://doi.org/10.24072/pci.animsci.100198.rev12








